Our research
The nucleosome is a dynamic epigenetic signaling hub for genome-templated processes. We use high-resolution structural biology, protein chemistry, and proteomics to understand the molecular mechanisms that govern epigenetic signaling in health and disease.
Determining the structural basis of histone modification cross-talk
By pairing atomic precision protein chemistry to build designer modified nucleosomes and high resolution structural biology, we aim to understand how signatures of epigenetic marks are established and how these signatures control recruitment of chromatin effectors that regulate gene expression, DNA replication, and DNA damage repair. Single particle cryo-electron microscopy (cryo-EM) and macromolecular X-ray crystallography allow us to visualize epigenetic enzymes in action on nucleosome substrates to uncover molecular mechanisms of signal integration on chromatin.
Establishing paradigms of nucleosome recognition
Despite decades of research, we still lack a fundamental understanding of how most epigenetic enzymes function in a chromatin environment. We are using proteomics to define patterns of nucleosome recognition with the goal of establishing a universal framework for nucleosome binding.
Creating tools to enable epigenetics research
We also are developing new tools to probe epigenetic signaling both in vitro and in model systems. Such tools will allow precise hypothesis-driven interrogation of epigenetic signaling networks.
Our Group




Recent News
08-24-2023 | Congratulations to Aleksandra on winning the UNC Postdoctoral Award for Research Excellence!

03-16-2023 | Congratulations to Cathy on UNC Dean’s Distinguished Dissertation Award!
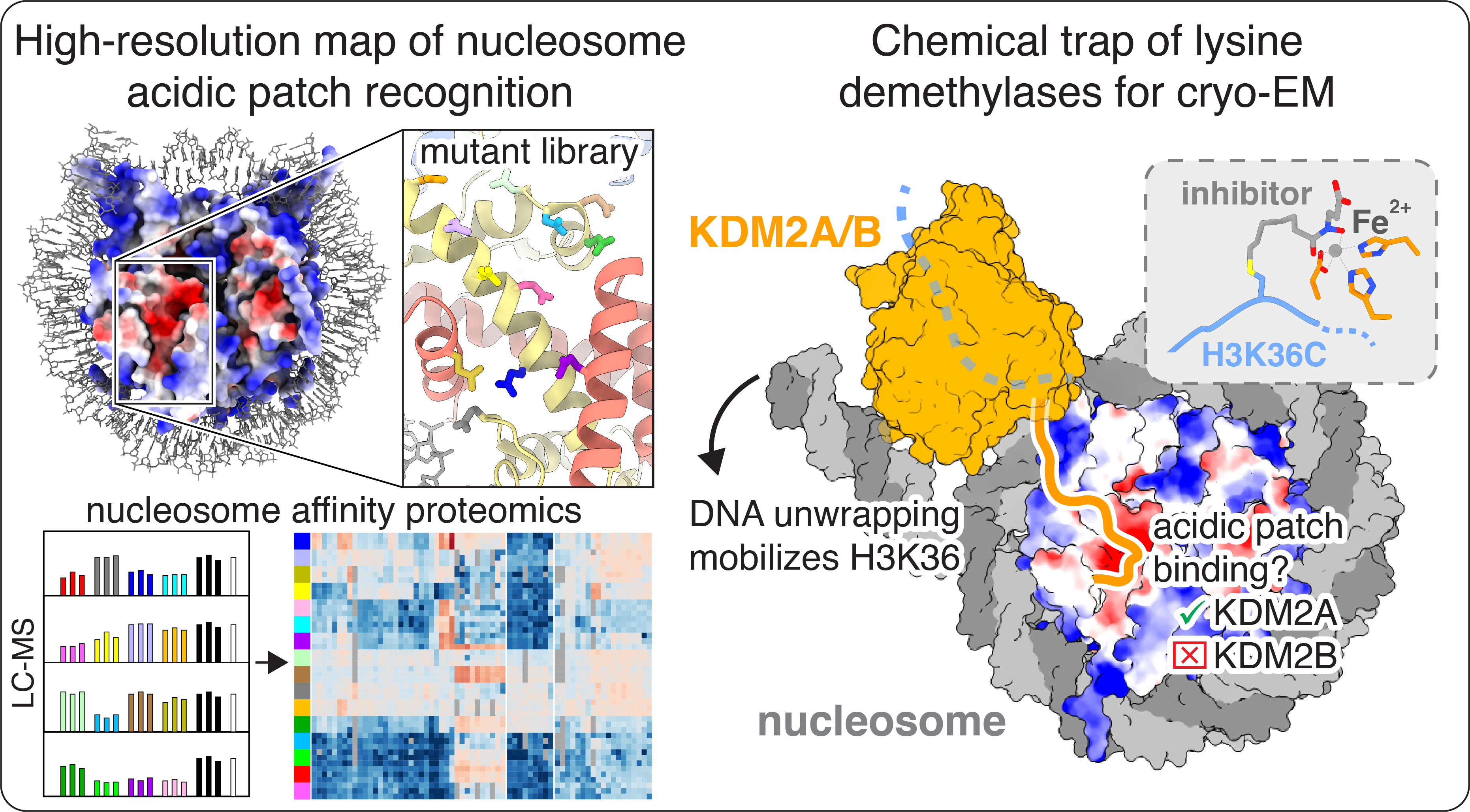
02-16-2023 | Cathy’s and Aleksandra’s paper is out today in Nature Chemical Biology!
10-31-2022 | Rob’s review in Current Opinion in Structural Biology is selected as runner-up for Best Paper of 2021!
08-17-2022 | Congratulations to Nate on successful defense of his PhD thesis!

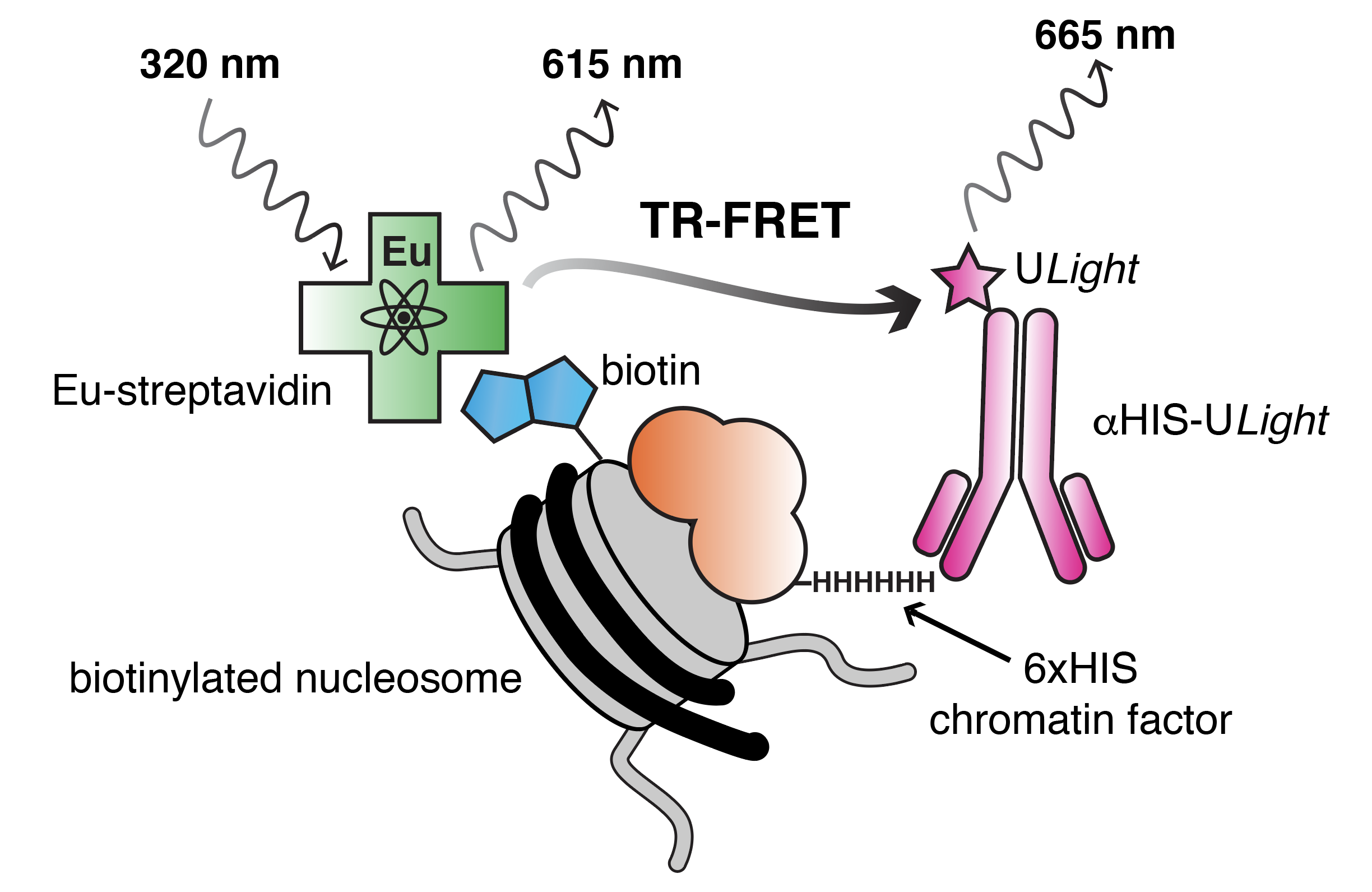
05-26-2022 | TR-FRET for nucleosome binding and foot-printing studies! Nate’s paper goes online today. Congrats Nate!
05-19-2022 | Gabby wins Diane Harris Leadership Award! A well-deserved honor for everything Gabby has done to promote diversity in STEM in the department and beyond. Congrats Gabby!

05-07-2022 | Congrats to Dalal on her graduation! A very successful undergrad tenure: three co-authored papers, an NSF GFRP, speaking at the Chancellor’s Science Scholars Finalist Weekend, and a Frances P. Venable Medal. Best of luck with your graduate studies at Duke!
03-23-2022 | Gabby defends thesis! An exciting semester for the McGinty lab.
